Chinese Medical Sciences Journal ›› 2019, Vol. 34 ›› Issue (3): 168-176.doi: 10.24920/003585
• Original Article • Previous Articles Next Articles
Identification of CDK6 and RHOU in Serum Exosome as Biomarkers for the Invasiveness of Non-functioning Pituitary Adenoma
Yu Shan1, Wang Xiaoshuang1, Cao Kaican2, Bao Xinjie3, *( ), Yu Jia1, *(
), Yu Jia1, *( )
)
- 1 State Key Laboratory of Medical Molecular Biology & Key Laboratory of RNA and Hematopoietic Regulation & Department of Biochemistry, Institute of Basic Medical Sciences, Chinese Academy of Medical Sciences & School of Basic Medicine, Peking Union Medical College, Beijing 100005, Chin; 2 Department of Thoracic Surgery, Nanfang Hospital, Guangzhou 510515, China;3 Department of Neurosurgery, Peking Union Medical College Hospital, Chinese Academy of Medical Sciences & Peking Union Medical College, Beijing 100730, Chin;
-
Received:2019-05-22Published:2019-09-30Online:2019-05-22 -
Contact:Bao Xinjie,Yu Jia E-mail:xinjieabao@163.com;j-yu@ibms.pumc.edu.cn
| In this article, the authors explored circulating biomarkers for screening the invasiveness of non-functioning pituitary adenomas, and found both CDK6 and RHOU showed significantly higher relative expression in invasive pituitary adenomas as compared to non-invasive pituitary adenomas, suggesting CDK6 and RHOU mRNA in serum exosomes can be used as markers for predicting invasiveness of pituitary adenomas, and combination of the two genes performs better. |
Cite this article
Yu Shan, Wang Xiaoshuang, Cao Kaican, Bao Xinjie, Yu Jia. Identification of CDK6 and RHOU in Serum Exosome as Biomarkers for the Invasiveness of Non-functioning Pituitary Adenoma[J].Chinese Medical Sciences Journal, 2019, 34(3): 168-176.
share this article
Add to citation manager EndNote|Reference Manager|ProCite|BibTeX|RefWorks
Table 1
Primers and probes for the target and reference genes"
| Gene name | Gene ID | Forward primer (5’-3’) | Reverse primer (5’-3’) | Probe (5’-3’) | Amplicon size (bp) |
|---|---|---|---|---|---|
| CDK6 | 1021 | CTAGCAACCATCCCTCCATTAC | CTCAGAGCATTCTGAAGACAGTAG | ACACAGAAAGCCCTCTTGAAGCAA | 104 |
| RHOU | 58480 | GCAGGGAGGTGAATACTCTTG | GAAGTACATCTTGGCCGACTTA | AAGGCCAACAGCAAGTGTTTGTGGGA | 92 |
| SPIRE2 | 84501 | GTCTGCACTTCCTGTAGCATAA | GATACCCTCTGAGGACTCTCAA | CACATCCCTGTCTACACACTGGGC | 98 |
| GAPDH | 2597 | GGTGTGAACCATGAGAAGTATGA | GAGTCCTTCCACGATACCAAAG | AGATCATCAGCAATGCCTCCTGCA | 123 |
Table 2
Clinicopathological features of NNF-PA and INF-PA patients included in the study"
| Groups | n | Age§ (yrs) | Gender (male/female, n) | p53 (n) | Ki-67 (n) | Largest diameter§ (mm) | Tumor texture (n) | ||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Negative | Positive | ≤3% | >3% | Hard | Soft | Other | |||||
| NNF | 32 | 47.28±16.30 | 13/19 | 30 | 2 | 27 | 5 | 21.19±7.68 | 9 | 21 | 2 |
| INF | 14 | 47.86±17.09 | 6/8 | 14 | 0 | 10 | 4 | 30.34±7.70 | 3 | 9 | 2 |
| t/χ2 value | 0.1095a | 0.0200b | 0.0292b | 0.3777b | 3.7152a | NA | |||||
| P value | >0.9999a | 0.8875b | 0.8644b | 0.5388b | 0.0006a | 0.7717c | |||||
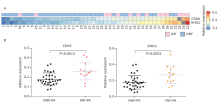

Figure 2.
Differential expression of CDK6 and RHOU in exosomes from NF-PAs. A. Heatmap of relative CDK6 and RHOU mRNA expression (normalized to GAPDH) in NNF-PAs and INF-PAs. Blocks filled with gray indicated the missing values. B. Statistical analysis of exosomal CDK6 and RHOU mRNA relative expression in NNF-PAs (n=32) compared with INF-PAs (n=13) samples. Data are shown as means±SD, compared with Unpaired t test."
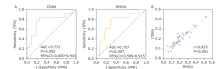
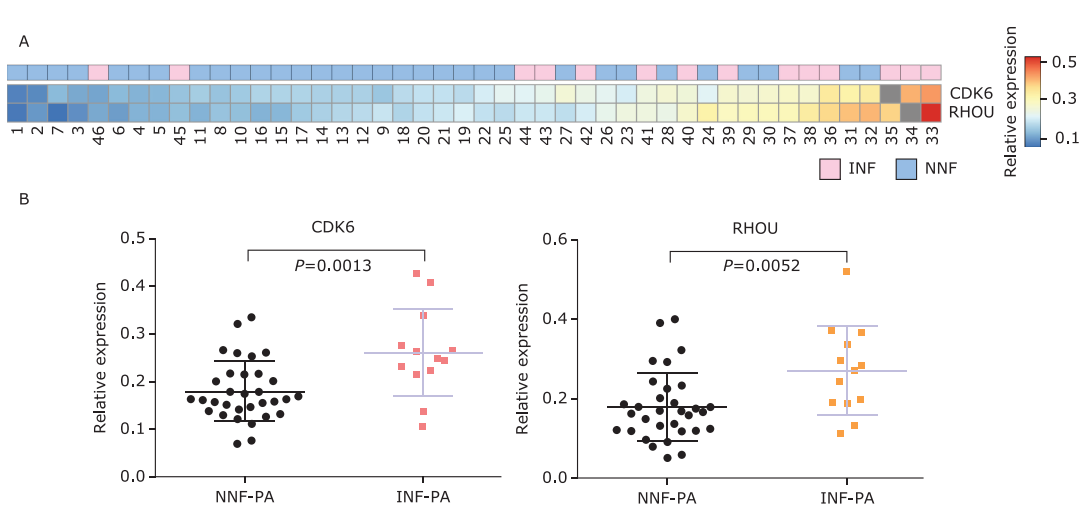

Figure 3.
The evaluation of performance using CDK6 or RHOU to discriminate INF-PAs from NNF-PAs. A. Receiver Operator Characteristic (ROC) curve of using CDK6 and RHOU exosomal mRNAs level to distinguish NNF-PAs from INF-PAs. B. The expressions of these two mRNAs in NF-PAs were prominently linear correlated."
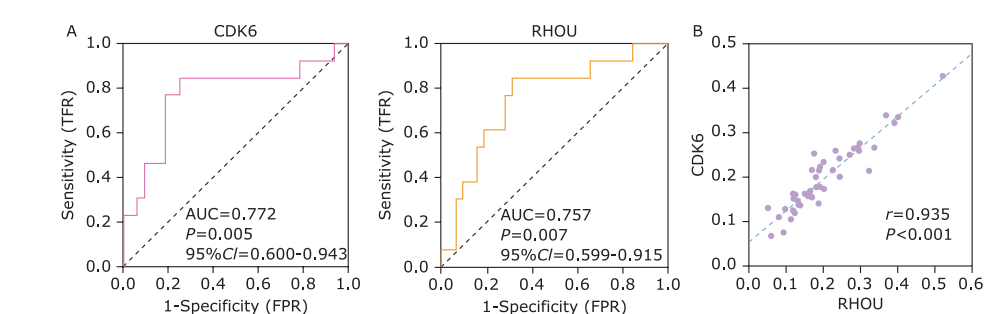
Table 3
Functional enrichments of CDK6 and RHOU"
| Items | Database | ID | Background number | Corrected P-value | Input (gene ID) |
|---|---|---|---|---|---|
| G1/S transition of mitotic cell cycle | Gene Ontology | GO:0000082 | 230 | 0.0071 | 58480|1021 |
| Cell cycle G1/S phase transition | Gene Ontology | GO:0044843 | 240 | 0.0071 | 58480|1021 |
| Mitotic cell cycle phase transition | Gene Ontology | GO:0044772 | 447 | 0.0136 | 58480|1021 |
| Cell cycle phase transition | Gene Ontology | GO:0044770 | 473 | 0.0136 | 58480|1021 |
| Mitotic cell cycle process | Gene Ontology | GO:1903047 | 867 | 0.0182 | 58480|1021 |
| Mitotic cell cycle | Gene Ontology | GO:0000278 | 937 | 0.0182 | 58480|1021 |
| Cell cycle process | Gene Ontology | GO:0022402 | 1299 | 0.0182 | 58480|1021 |
| Cytoskeletal part | Gene Ontology | GO:0044430 | 1468 | 0.0182 | 58480|1021 |
| Cell cycle | Gene Ontology | GO:0007049 | 1665 | 0.0182 | 58480|1021 |
| Cell projection | Gene Ontology | GO:0042995 | 1772 | 0.0182 | 58480|1021 |
| Purine ribonucleoside triphosphate binding | Gene Ontology | GO:0035639 | 1788 | 0.0182 | 58480|1021 |
| Purine ribonucleoside binding | Gene Ontology | GO:0032550 | 1798 | 0.0182 | 58480|1021 |
| Ribonucleoside binding | Gene Ontology | GO:0032549 | 1801 | 0.0182 | 58480|1021 |
| Purine nucleoside binding | Gene Ontology | GO:0001883 | 1801 | 0.0182 | 58480|1021 |
| Nucleoside binding | Gene Ontology | GO:0001882 | 1808 | 0.0182 | 58480|1021 |
| Purine ribonucleotide binding | Gene Ontology | GO:0032555 | 1830 | 0.0182 | 58480|1021 |
| Purine nucleotide binding | Gene Ontology | GO:0017076 | 1843 | 0.0182 | 58480|1021 |
| Ribonucleotide binding | Gene Ontology | GO:0032553 | 1846 | 0.0182 | 58480|1021 |
| Cytoskeleton | Gene Ontology | GO:0005856 | 1983 | 0.0197 | 58480|1021 |
Table 4
Comparisons of the performance using CDK6, RHOU or both to distinguish between NNF-PAs and INF-PAs"
| Items | CDK6 | RHOU | Combination |
|---|---|---|---|
| Total | 44 | 44 | 44 |
| Positives (P, n) | 12 | 12 | 12 |
| Negatives (N, n) | 32 | 32 | 32 |
| True positives (TP, n) | 10 | 10 | 10 |
| False positives (FP, n) | 8 | 10 | 6 |
| False negatives (FN, n) | 2 | 2 | 2 |
| True negatives (TN, n) | 24 | 22 | 26 |
| Accuracy (%) | 77.27 | 72.73 | 81.82 |
| Sensitivity (%) | 83.33 | 83.33 | 83.33 |
| Specificity (%) | 75.00 | 68.75 | 81.25 |
| Precision (%) | 55.56 | 50.00 | 62.50 |
| 1. | Asa SL, Ezzat S . The cytogenesis and pathogenesis of pituitary adenomas. Endocr Rev 1998; 19(6):798-827. doi: 10.1210/edrv.19.6.0350. |
| 2. | Daly AF, Rixhon M, Adam C , et al. High prevalence of pituitary adenomas: a cross-sectional study in the province of Liege, Belgium. J Clin Endocrinol Metab 2006; 91(12):4769-75. doi: 10.1210/jc.2006-1668. |
| 3. | Fernandez A, Karavitaki N, Wass JA . Prevalence of pituitary adenomas: a community-based, cross-sectional study in Banbury (Oxfordshire, UK). Clin Endocrinol (Oxf) 2010; 72(3):377-82. doi: 10.1111/j.1365-2265.2009.03667.x. |
| 4. | Lopes MBS . The 2017 World Health Organization classification of tumors of the pituitary gland: a summary. Acta Neuropathol 2017; 134(4):521-35. doi: 10.1007/s00401-017-1769-8. |
| 5. | Selman WR, Laws ER, Jr., Scheithauer BW , et al. The occurrence of dural invasion in pituitary adenomas. J Neurosurg 1986; 64(3):402-7. doi: 10.3171/jns.1986.64.3.0402. |
| 6. | Thapar K, Kovacs K, Scheithauer BW , et al. Proliferative activity and invasiveness among pituitary adenomas and carcinomas: an analysis using the MIB-1 antibody. Neurosurgery 1996; 38(1):99-106; discussion 106-7. doi: 10.1097/00006123-199601000-00024. |
| 7. | Raverot G, Burman P, McCormack A , et al. European Society of Endocrinology Clinical Practice Guidelines for the management of aggressive pituitary tumours and carcinomas. Eur J Endocrinol 2018; 178(1):G1-g24. doi: 10.1530/eje-17-0796. |
| 8. | Boriachek K, Islam MN, Moller A , et al. Biological functions and current advances in isolation and detection strategies for exosome nanovesicles. Small 2018; 14(6). doi: 10.1002/smll.201702153. |
| 9. | Vlassov AV, Magdaleno S, Setterquist R , et al. Exosomes: current knowledge of their composition, biological functions, and diagnostic and therapeutic potentials. Biochim Biophys Acta 2012; 1820(7):940-8. doi: 10.1016/j.bbagen.2012.03.017. |
| 10. | Del Re M, Biasco E, Crucitta S , et al. The detection of androgen receptor splice variant 7 in plasma-derived exosomal RNA strongly predicts resistance to hormonal therapy in metastatic prostate cancer patients. Eur Urol 2017; 71(4):680-7. doi: 10.1016/j.eururo.2016.08.012. |
| 11. | Knosp E, Steiner E, Kitz K , et al. Pituitary adenomas with invasion of the cavernous sinus space: a magnetic resonance imaging classification compared with surgical findings. Neurosurgery 1993; 33(4):610-7; discussion 7-8. doi: 10.1097/00006123-199310000-00008 |
| 12. | Corbisier P, Pinheiro L, Mazoua S , et al. DNA copy number concentration measured by digital and droplet digital quantitative PCR using certified reference materials. Anal Bioanal Chem 2015; 407(7):1831-40. doi: 10.1007/s00216-015-8458-z. |
| 13. | Asa SL, Ezzat S . The pathogenesis of pituitary tumors. Annu Rev Pathol 2009; 4(1):97-126. doi: 10.1146/annurev.pathol.4.110807.092259. |
| 14. | Sav A, Rotondo F, Syro LV , et al. Biomarkers of pituitary neoplasms. Anticancer Res 2012; 32(11):4639-54. |
| 15. | Amar AP, Hinton DR, Krieger MD , et al. Invasive pituitary adenomas: significance of proliferation parameters. Pituitary 1999; 2(2):117-22. |
| 16. | Knosp E, Kitz K, Perneczky A . Proliferation activity in pituitary adenomas: measurement by monoclonal antibody Ki-67. Neurosurgery 1989; 25(6):927-30. doi: 10.1227/00006123-198912000-00012 |
| 17. | Zhao X, Li J, Huang S , et al. MiRNA-29c regulates cell growth and invasion by targeting CDK6 in bladder cancer. Am J Transl Res 2015; 7(8):1382-9. |
| 18. | Mendrzyk F, Radlwimmer B, Joos S , et al. Genomic and protein expression profiling identifies CDK6 as novel independent prognostic marker in medulloblastoma. J Clin Oncol 2005; 23(34):8853-62. doi: 10.1200/jco.2005.02.8589. |
| 19. | Costello JF, Plass C, Arap W , et al. Cyclin-dependent kinase 6 (CDK6) amplification in human gliomas identified using two-dimensional separation of genomic DNA. Cancer Res 1997; 57(7):1250-4. |
| 20. | Bax DA, Mackay A, Little SE , et al. A distinct spectrum of copy number aberrations in pediatric high-grade gliomas. Clin Cancer Res 2010; 16(13):3368-77. doi: 10.1158/1078-0432.Ccr-10-0438. |
| 21. | Nagel S, Leich E, Quentmeier H , et al. Amplification at 7q22 targets cyclin-dependent kinase 6 in T-cell lymphoma. Leukemia 2008; 22(2):387-92. doi: 10.1038/sj.leu.2405028. |
| 22. | Nebenfuehr S, Bellutti F, Sexl V . Cdk6: at the interface of Rb and p53. Mol Cell Oncol 2018; 5(5):e1511206. doi: 10.1080/23723556.2018.1511206. |
| 23. | Lee KH, Lotterman C, Karikari C , et al. Epigenetic silencing of MicroRNA miR-107 regulates cyclin-dependent kinase 6 expression in pancreatic cancer. Pancreatology 2009; 9(3):293-301. doi: 10.1159/000186051. |
| 24. | Lim JT, Mansukhani M, Weinstein IB . Cyclin-dependent kinase 6 associates with the androgen receptor and enhances its transcriptional activity in prostate cancer cells. Proc Natl Acad Sci U S A 2005; 102(14):5156-61. doi: 10.1073/pnas.0501203102. |
| 25. |
Wang G, Zheng L, Yu Z , et al. Increased cyclin-dependent kinase 6 expression in bladder cancer. Oncol Lett 2012; 4(1):43-6. doi: 10.3892/ol.2012.695.
doi: 10.3892/ol.2012.695 |
| 26. | Symons M, Segall JE . Rac and Rho driving tumor invasion: who’s at the wheel? Genome Biol 2009; 10(3):213. doi: 10.1186/gb-2009-10-3-213. |
| 27. | Liu S, Wang Y, Xue W , et al. Genetic variants in the genes encoding rho GTPases and related regulators predict cutaneous melanoma-specific survival. Int J Cancer 2017; 141(4):721-30. doi: 10.1002/ijc.30785. |
| 28. | Vega FM, Ridley AJ . Rho GTPases in cancer cell biology. FEBS Lett 2008; 582(14):2093-101. doi: 10.1016/j.febslet.2008.04.039. |
| 29. | Parri M, Chiarugi P . Rac and Rho GTPases in cancer cell motility control. Cell Commun Signal 2010; 8(1):23. doi: 10.1186/1478-811x-8-23. |
| 30. | Simpson KJ, Dugan AS, Mercurio AM . Functional analysis of the contribution of RhoA and RhoC GTPases to invasive breast carcinoma. Cancer Res 2004; 64(23):8694-701. doi: 10.1158/0008-5472.Can-04-2247. |
| 31. | Canovas Nunes S, Manzoni M, Pizzi M , et al. The small GTPase RhoU lays downstream of JAK/STAT signaling and mediates cell migration in multiple myeloma. Blood Cancer J 2018; 8(2):20. doi: 10.1038/s41408-018-0053-z. |
| 32. |
Bhavsar PJ, Infante E, Khwaja A , et al. Analysis of Rho GTPase expression in T-ALL identifies RhoU as a target for Notch involved in T-ALL cell migration. Oncogene 2013; 32(2):198-208. doi: 10.1038/onc.2012.42.
doi: 10.1038/onc.2012.42 |
| 33. | Dart AE, Box GM, Court W , et al. PAK4 promotes kinase-independent stabilization of RhoU to modulate cell adhesion. J Cell Biol 2015; 211(4):863-79. doi: 10.1083/jcb.201501072. |
| 34. | Hodge RG, Ridley AJ . Regulation and functions of RhoU and RhoV. Small GTPases 2017; 30:1-8. doi: 10.1080/21541248.2017.1362495. |
| 35. | Faure S, Fort P . Atypical RhoV and RhoU GTPases control development of the neural crest. Small GTPases 2015; 6(4):174-7. doi: 10.1080/21541248.2015.1025943. |
| [1] | Wei Yang, Mei Zhang. Biomarkers with Potential Predictive Value for Cardiotoxicity in Anticancer Treatments [J]. Chinese Medical Sciences Journal, 2021, 36(4): 333-341. |
| [2] | Yi Wang, Zhuhua Zhang, Wei Lian. Sudden Sensorineural Hearing Loss after Pituitary Adenoma Resection—A Case Series with Literature Review [J]. Chinese Medical Sciences Journal, 2021, 36(2): 120-126. |
| [3] | Weijia Wang, Le Shen, Labaciren , Hange Li, Yuelun Zhang, Yuguang Huang. Evaluation of Burnout Among Anesthesiologists Working in Tibet, China: Altitude and Attitude [J]. Chinese Medical Sciences Journal, 2021, 36(2): 97-102. |
| [4] | Wang Zhan, Wang Xu, Wang Wenda, Zheng Guoyang, Guo Hao, Zhang Yushi. Value of Preoperative Neutrophil-to-Lymphocyte Ratio in Predicting Prognosis of Surgically Resectable Urinary Cancers: Systematic Review and Meta-Analysis [J]. Chinese Medical Sciences Journal, 2020, 35(3): 262-271. |
| [5] | Wu Jianqiang, Qin Weiwei, Pan Li, Wang Xiaorong, Zhang Biao, Shan Guangliang, Gao Youhe. Regional Differences of the Urinary Proteomes in Healthy Chinese Individuals [J]. Chinese Medical Sciences Journal, 2019, 34(3): 157-167. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
|